Bioinformatics Toolbox0 pages
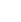
Bioinformatics Toolbox
Read, analyze, and visualize genomic and proteomic data
Bioinformatics Toolbox™ provides algorithms and visualization techniques for Next Generation Sequencing
(NGS), microarray analysis, mass spectrometry, and gene ontology. Using toolbox functions, you can read
genomic and proteomic data from standard file formats such as SAM, FASTA, CEL, and CDF, as well as from
online databases such as the NCBI Gene Expression Omnibus and GenBank®. You can explore and visualize this
data with sequence browsers, spatial heatmaps, and clustergrams. The toolbox also provides statistical techniques
for detecting peaks, imputing values for missing data, and selecting features.
You can combine toolbox functions to support common bioinformatics workflows. You can use ChIP-Seq data to
identify transcription factors; analyze RNA-Seq data to identify differentially expressed genes; identify copy
number variants and SNPs in microarray data; and classify protein profiles using mass spectrometry data.
Learn more about computational biology.
Key Features
▪ Next Generation Sequencing analysis and browser
▪ Sequence analysis and visualization, including pairwise and multiple sequence alignment and peak detection
▪ Microarray data analysis, including reading, filtering, normalizing, and visualization
▪ Mass spectrometry analysis, including preprocessing, classification, and marker identification
▪ Phylogenetic tree analysis
▪ Graph theory functions, including interaction maps, hierarchy plots, and pathways
▪ Data import from genomic, proteomic, and gene expression files, including SAM, FASTA, CEL, and CDF,
and from databases such as NCBI and GenBank
What additional features would you like in Bioinformatics Toolbox?
1
"
 عضویت
عضویت  ورود اعضا
ورود اعضا راهنمای خرید
راهنمای خرید